Software

BASiCS
BASiCS (Bayesian Analysis of Single-Cell Sequencing data) is an integrated Bayesian hierarchical model to perform statistical analyses of single-cell RNA sequencing datasets in the context of supervised experiments (where the groups of cells of interest are known a priori, e.g. experimental conditions or cell types). BASiCS performs built-in data normalisation (global scaling) and technical noise quantification. BASiCS also provides probabilistic frameworks for (i) highly (or lowly) variable genes within a single group of cells and (ii) differential expression analyses (mean and variability) between groups of cells.
Category: scRNA-seq, Bayesian Modelling
Language: R


BASSLINE
Mixtures of life distributions provide a convenient framework for survival analysis: particularly when standard models such as the Weibull or the log-normal are unable to capture some features from the data. These mixtures can also account for unobserved heterogeneity or outlying observations. BASSLINE (BAyeSian Survival anaLysIs usiNg shapE mixtures of log-normal distributions) uses shape mixtures of log-normal distributions to fit data with fat tails and has been adapted from code written by Vallejos and Steel. Some of the functions have been rewritten in C++ for increased performance.
Category: Survival Analysis
Language: R
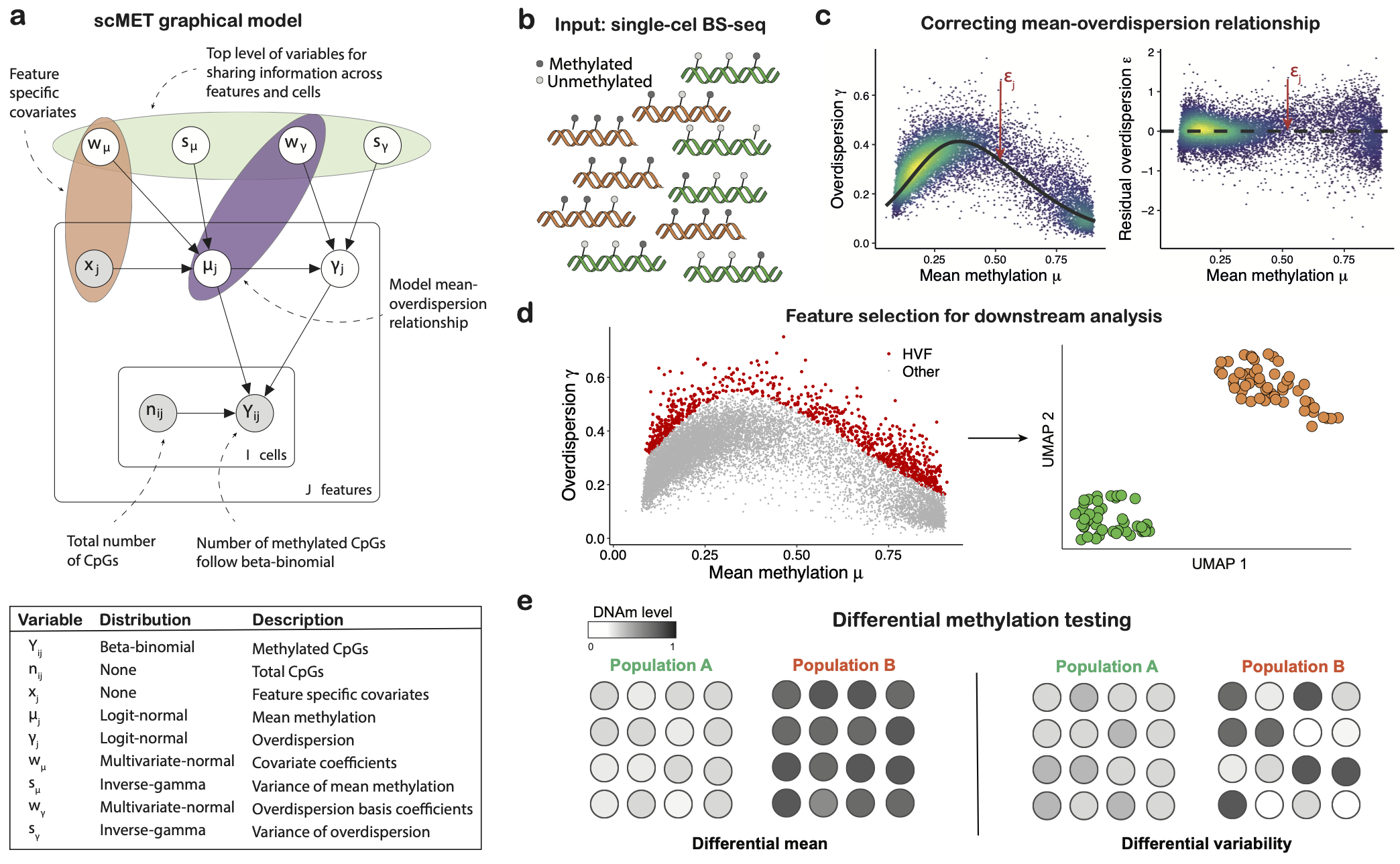
scMET
A Bayesian framework for the analysis of single cell DNA methylation data. This modelling approach combines a hierarchical beta-binomial specification with a generalised linear model framework with the aim of capturing biological overdispersion and overcome data sparsity by sharing information across cells and genomic features.
Category: scDNAm, Bayesian Modelling
Language: R